-Search query
-Search result
Showing all 50 items for (author: sherpa & d)

EMDB-19039: 
Map of YPEL5-bound WDR26 dimer obtained by focused refinement of the WDR26-CTLH subcomplex
Method: single particle / : Chrustowicz J, Schulman BA

EMDB-17705: 
Structure of Chelator-GIDSR4 - Fbp1 - phospho-Ubc8~ubiquitin - class I
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-17706: 
Structure of Chelator-GIDSR4 - Fbp1 - phospho-Ubc8~ubiquitin - class II
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-17707: 
Chelator-GIDSR4 - Fbp1 - phospho-Ubc8~ubiquitin - class III
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-17709: 
Structure of Chelator-GIDSR4 - Fbp1 - phospho-Ubc8~ubiquitin - class V
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-17710: 
Structure of Chelator-GIDSR4 - Fbp1 - phospho-Ubc8~ubiquitin - class IV
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-17713: 
Catalytic module of human CTLH E3 ligase bound to multiphosphorylated UBE2H~ubiquitin
Method: single particle / : Chrustowicz J, Sherpa D, Prabu RJ, Schulman BA

EMDB-17715: 
SRS and Cat modules of human CTLHSR4 bound to multiphosphorylated UBE2H~ubiquitin
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-17716: 
Structure of CTLHSR4 - phospho-UBE2H~ubiquitin bound to engineered VH
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA
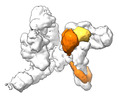
EMDB-17717: 
SRS and Cat modules of yeast Chelator-GIDSR4 bound to multiphosphorylated Ubc8~ubiquitin
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA
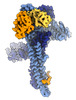
EMDB-17764: 
Catalytic module of yeast GID E3 ligase bound to multiphosphorylated Ubc8~ubiquitin
Method: single particle / : Chrustowicz J, Sherpa D, Prabu RJ, Schulman BA

PDB-8pjn: 
Catalytic module of human CTLH E3 ligase bound to multiphosphorylated UBE2H~ubiquitin
Method: single particle / : Chrustowicz J, Sherpa D, Prabu RJ, Schulman BA

PDB-8pmq: 
Catalytic module of yeast GID E3 ligase bound to multiphosphorylated Ubc8~ubiquitin
Method: single particle / : Chrustowicz J, Sherpa D, Prabu RJ, Schulman BA

EMDB-16242: 
Cryo-EM structure of RANBP10-CTLH SR4 complex
Method: single particle / : Sherpa D, Chrustowicz J

EMDB-16243: 
Cryo-EM map of ARMC8-specific nanobody bound to CTLH-SR4
Method: single particle / : Chrustowicz J, Sherpa D

EMDB-14323: 
Structure of Chelator-GIDSR4 bound to Mdh2
Method: single particle / : Sherpa D, Chrustowicz J, Schulman B

EMDB-14324: 
Structure of Cage-GIDSR4 bound to PHSVTP-Fbp1
Method: single particle / : Chrustowicz J, Sherpa D, Qiao S, Schulman B

EMDB-14338: 
Structure of endogenous Cage-GIDAnt complex
Method: single particle / : Sherpa D, Chrustowicz J, Qiao S, Schulman B

EMDB-32830: 
GID subcomplex: Gid12 bound Substrate Receptor Scaffolding module
Method: single particle / : Qiao S, Cheng JD, Schulman BA

EMDB-32831: 
Gid12 bound GIDSR4 E3 ubiquitin ligase complex
Method: single particle / : Qiao S, Cheng DJ, Schulman BA
EMDB-32833: 
Gid12 bound Chelator-GIDSR4
Method: single particle / : Qiao S, Cheng DJ, Schulman BA

EMDB-32834: 
Cage assembly GID E3 ubiquitin ligase
Method: single particle / : Qiao S, Cheng DJ, Schulman BA

EMDB-32835: 
Gid12 bound Cage-GIDSR3
Method: single particle / : Qiao S, Cheng DJ, Schulman BA

PDB-7wug: 
GID subcomplex: Gid12 bound Substrate Receptor Scaffolding module
Method: single particle / : Qiao S, Cheng JD, Schulman BA

EMDB-12537: 
Structure of human CTLH SR4 complex
Method: single particle / : Sherpa D, Chrustowicz J, Schulman BA

EMDB-12538: 
Structure of endogenous yeast GID Ant complex
Method: single particle / : Sherpa D, Chrustowicz J, Qiao S, Schulman BA

EMDB-12540: 
Structure of endogenous yeast Chelator-GID Ant complex
Method: single particle / : Sherpa D, Chrustowicz J, Qiao S, Schulman BA

EMDB-12541: 
Structure of Apo Chelator-GID SR4 E3 ubiquitin ligase
Method: single particle / : Sherpa D, Chrustowicz J, Qiao S, Prabu JR, Schulman BA

EMDB-12542: 
Structure of human CTLH-WDR26 supramolecular assembly
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-12545: 
Structure of supramolecular assembly and substrate receptor scaffolding modules of human CTLH-WDR26 complex
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-12547: 
Structure of supramolecular assembly and substrate receptor scaffolding modules of human CTLH-MKLN1 complex
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-12548: 
Structure of GID SR4 complex
Method: single particle / : Sherpa D, Chrustowicz J, Schulman BA

EMDB-12557: 
Structure of yeast Chelator-GID SR4 E3 ubiquitin ligase bound to its tetrameric metabolic enzyme substrate Fbp1
Method: single particle / : Chrustowicz J, Sherpa D, Prabu JR, Schulman BA

EMDB-12559: 
Substrate receptor scaffolding module of yeast Chelator-GID SR4 E3 ubiquitin ligase bound to Fbp1 substrate
Method: single particle / : Sherpa D, Chrustowicz J, Prabu JR, Schulman BA

EMDB-12560: 
Catalytic module of yeast Chelator-GID SR4 E3 ubiquitin ligase
Method: single particle / : Sherpa D, Chrustowicz J, Prabu JR, Schulman BA

EMDB-12563: 
Supramolecular assembly module of yeast Chelator-GID SR4 E3 ubiquitin ligase
Method: single particle / : Chrustowicz J, Sherpa D, Prabu JR, Schulman BA

EMDB-12564: 
Substrate receptor scaffolding module of human CTLH E3 ubiquitin ligase
Method: single particle / : Chrustowicz J, Sherpa D, Prabu JR, Schulman BA

PDB-7ns3: 
Substrate receptor scaffolding module of yeast Chelator-GID SR4 E3 ubiquitin ligase bound to Fbp1 substrate
Method: single particle / : Sherpa D, Chrustowicz J, Prabu JR, Schulman BA

PDB-7ns4: 
Catalytic module of yeast Chelator-GID SR4 E3 ubiquitin ligase
Method: single particle / : Sherpa D, Chrustowicz J, Prabu JR, Schulman BA

PDB-7nsb: 
Supramolecular assembly module of yeast Chelator-GID SR4 E3 ubiquitin ligase
Method: single particle / : Chrustowicz J, Sherpa D, Prabu JR, Schulman BA

PDB-7nsc: 
Substrate receptor scaffolding module of human CTLH E3 ubiquitin ligase
Method: single particle / : Chrustowicz J, Sherpa D, Prabu JR, Schulman BA

EMDB-10326: 
Structure of inactive GID E3 ubiquitin ligase complex
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-10327: 
Structure of active GID E3 ubiquitin ligase complex
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-10328: 
Structure of GID Scaffold subcomplex
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-10329: 
Structure of GID Scaffold subcomplex bound to substrate receptor Gid10
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-10330: 
Structure of GID Scaffold Subcomplex bound to substrate receptor Gid4
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-10331: 
Structure of endogenous inactive GID E3 ubiquitin ligase complex
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-10332: 
Structure of active GID E3 ubiquitin ligase complex with RING domains deletion
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-10333: 
Structure of active GID E3 ubiquitin ligase complex minus Gid2 and delta Gid9 RING domain
Method: single particle / : Qiao S, Prabu JR, Schulman BA

PDB-6swy: 
Structure of active GID E3 ubiquitin ligase complex minus Gid2 and delta Gid9 RING domain
Method: single particle / : Qiao S, Prabu JR, Schulman BA
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
